We Study the Host-Pathogen Kinetics of Influenza Virus Infections and Bacterial Coinfection
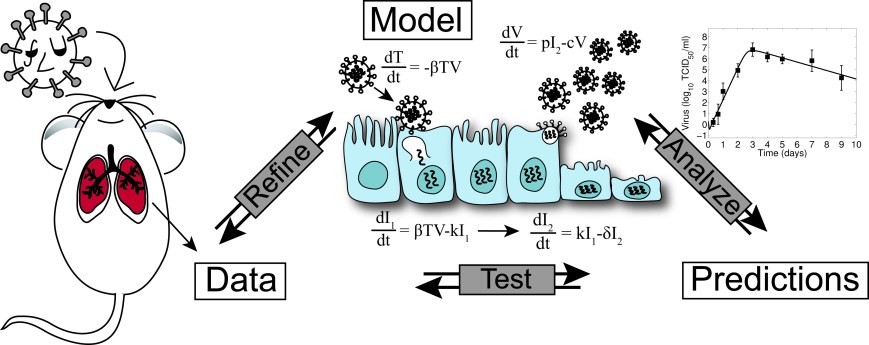
Our Approach: We operate at the interface of mathematics and biology. We use an integrative approach consisting of data-driven mathematical models together with model-driven experiments to study in vivo host-pathogen and pathogen-pathogen dynamics.
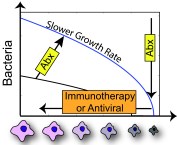
Why Study Coinfections? Influenza viruses pose a considerable threat to public health each year during seasonal epidemics and even more so when a pandemic strain emerges. Bacterial co-pathogens often complicate influenza virus infections and have accounted for ~40-95% of influenza-associated mortality in past pandemics. There are numerous host and pathogen factors that contribute to infection and coinfection pathogenicity. Understanding the relative contribution and time scale of each is difficult, which makes developing effective treatments challenging.
Why Use Mathematics? Mathematical models provide a rigorous way to extract information about infection processes, determine the time scales of these processes and how can be perturbed with therapeutics, simultaneously evaluate potential disease mechanisms, and develop new hypotheses about the underlying biology. Models are particularly useful to tease apart the complexities of multi-pathogen infections. We build and utilize ordinary differential equation models and analysis, model selection theory, perturbation theory, data fitting and parameter estimation.
Why Validate Theory With Experiments? The predictions that come from mathematical analyses are typically accurate but often go untested. Further, many models use approximations to make predictions but lack the data necessary to increase the mechanistic detail. We have changed this paradigm by designing and carrying out experiments that test specific analytical predictions. By doing so, we gain additional biological insight and are able to increase the detail of our models. See our publication page for details.
Projects
|
Viral Kinetics
|
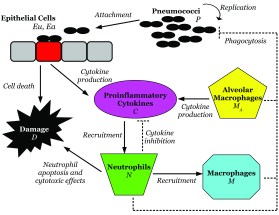
Mechanisms of Coinfection
|
|
|
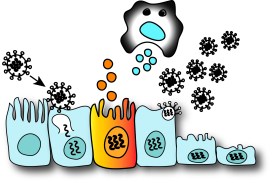
Immune Response Kinetics |
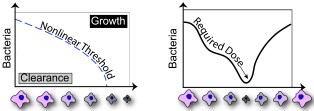
Bacterial Kinetics |
|
|
Treatment Strategies |
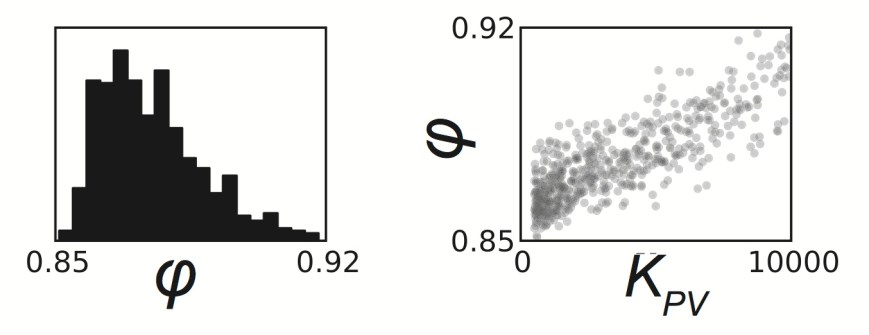
Parameter Estimation
|